The use and applicability of Internet search queries for infectious disease surveillance in low- to middle-income countries
Abstract
Uncontrolled outbreaks of emerging infectious diseases can pose threats to livelihoods and can undo years of progress made in developing regions, such as Sub-Saharan Africa. Therefore, the surveillance and early outbreak detection of infectious diseases, e.g., Dengue fever, is crucial. As a low-cost and timely source, Internet search queries data [e.g., Google Trends data (GTD)] are used and applied in epidemiological surveillance. This review aims to identify and evaluate relevant studies that used GTD in prediction models for epidemiological surveillance purposes regarding emerging infectious diseases. A comprehensive literature search in PubMed/MEDLINE was carried out, using relevant keywords identified from up-to-date literature and restricted to low- to middle-income countries. Eight studies were identified and included in the current review. Three focused on Dengue fever, three analyzed Zika virus infections, and two were about COVID-19. All studies investigated the correlation between GTD and the cases of the respective infectious disease; five studies used additional (time series) regression analyses to investigate the temporal relation. Overall, the reported positive correlations were high for Zika virus (0.75-0.99) or Dengue fever (0.87-0.94) with GTD, but not for COVID-19 (-0.81 to 0.003). Although the use of GTD appeared effective for infectious disease surveillance in low- to middle-income countries, further research is needed. The low costs and availability remain promising for future surveillance systems in low- to middle-income countries, but there is an urgent need for a standard methodological framework for the use and application of GTD.
Keywords
INTRODUCTION
Globally, the number of different emerging infectious diseases is increasing[1]. They are defined as “diseases that are newly recognized in a population or have existed but are rapidly increasing in incidence or geographic range”[2].
According to the World Health Organization (WHO)[3], in low-income countries, communicable neonatal conditions and lower respiratory infections are common causes of death. In lower- to middle-income countries, communicable neonatal conditions and lower-respiratory infections each caused over one million deaths in 2019 and remain the third and fifth leading causes of death worldwide, respectively[3].
In areas with inadequate capacity to detect and monitor local disease incidence, infectious diseases often spread unnoticed, whereas detecting disease outbreaks early and taking targeted countermeasures is a challenge for national health systems. This is particularly faced by developing regions, such as countries in Sub-Saharan Africa[4].
Outbreaks can also pose threats to economies and livelihoods and undo decades of progress. They represent a major public health concern[1]. Therefore, the prevention of uncontrolled outbreaks is fundamental. The more quickly disease outbreaks are recognized, the earlier investigations and control measures can be applied. Hence, surveillance systems are used to monitor timely information for outbreak detection or characterization[5].
Apart from traditional surveillance approaches, when routine health data are available and accessible, “syndromic surveillance” tools have been incorporated to “perform statistical tests for aberrant temporal or temporal/geographical trends” [5] so that epidemiologists and public health experts can assess the results and implement countermeasures. Syndromic surveillance uses the disease syndromes, e.g., symptoms or other indicators of the particular disease, and is a useful application if the cause of the disease or the outbreak is yet unknown. Syndromic surveillance aims to rapidly monitor syndromic categories of diseases, usually daily or more frequently[5]. Nevertheless, the use of health records is expensive, and therefore its use for timely outbreak detection is limited[6]. In any case, information searching is a human trait.
In the era of digitalization, searching the Internet is the first address when somebody has a question about a certain topic. Half of the assessed participants in a cross-sectional study searched online for information before they consulted their general practitioner[7]. In evolving situations, such as during an epidemic, researchers try to utilize this information-seeking behavior of a population by using the Internet search queries to predict potential outbreaks[8].
In 2020, the most searched term worldwide was “coronavirus”[9]. This “trend” is summarized by Google to show the change in online interests in time series for a selected period and region. The data, which are normalized by Google into the relative search volumes (RSVs), can be downloaded directly from the Web in a comma-separated value format. For this reason, different studies used Google trends data (GTD) to model the spread of the coronavirus in selected regions[10-12]. However, the principle to use GTD for epidemiological surveillance was not a novel idea.
Already in 2008, Google introduced flu trends to predict doctor visits for influenza-like illness in the US[13]. The motivation was to supplement traditional surveillance systems to help to predict outbreaks of the flu[14]. In 2013, the GTD model predicted more than double the proportion of physician consultation than the center for disease control and prevention (CDC)[15]. The model was overfitted, a phenomenon when prediction models, algorithms, and computational techniques are too specific[1]. Therefore, the regression line describing the model traverses each data point, “finding” random patterns in the training data that are not part of the true model in the population it is generalized to[16]. Google stopped the Flu Trends project in 2015; Google Dengue Trends (2004-2015) is also no longer available.
In 2009, Gunther Eysenbach introduced the terms “infodemiology” and “infoveillance”[17]. The terms combine the word information and epidemiology or surveillance to demonstrate the symbiosis of the sciences. He defined “infodemiology” as the “science of distribution and determinants of information in an electronic medium, specifically the Internet, or in a population, with the ultimate aim to inform public health and public policy”[17]. The corresponding use of infodemiology data was therefore called “infoveillance”[17].
Since the introduction of the terms, Internet data have been increasingly integrated into epidemiological research[18]. However, it is not yet known in which settings the use of GTD in epidemiological surveillance for emerging infectious diseases gives valid and reliable results. The literature suggests that unlimited Internet access is crucial to receive data that represent the information-seeking behavior of the whole population[18].
A growing digital infrastructure, particularly in the mobile sector in regions of Sub-Saharan Africa, and technological advances offer promising opportunities for more efficient information and knowledge transfer to improve epidemiological health surveillance. The collection of information from mobile devices and the use of open data sources, e.g., freely accessible population and environmental data, hold enormous potential as a new component of traditional surveillance systems, which at the same time requires intelligent networking of qualified actors across national borders and individual disciplines.
To receive relevant information on the use and applicability of GTD in epidemiological surveillance in other low- to middle-income regions in the world, this review aims to identify relevant studies that used GTD in prediction models for epidemiological surveillance purposes regarding emerging infectious diseases such as HIV infections, SARS, COVID-19, Lyme disease, Dengue fever, E. coli, Hantavirus, West Nile virus, or Zika virus infection[19].
METHODS
Search strategy
Relevant keywords were identified from the articles by Choi et al.[20], Barros et al.[21], and Milinovich et al.[6] . The applied string of keywords is listed below in Table 1. The searched database was Medline through PubMed. There were no restrictions regarding the calendar date, and no filters were applied. Only publications in English were considered.
Search strategy
| Search strategy item | Search strategy details |
| Searched database | PubMed/MEDLINE |
| Used string of keywords | (“surveillance systems”[All Fields]) OR (“syndromic”[All Fields] AND “surveillance”[All Fields]) OR ((“digital”[All Fields] OR “early”[All Fields]) AND “detection”[All Fields]) OR “biosurveillance”[MeSH Terms] OR “infoveillance”[All Fields] OR “infodemiology”[All Fields] OR (“online”[All Fields] AND “surveillance”[All Fields]) OR (“outbreaks”[All Fields] AND “forecasting”[MeSH Terms]) OR (“web”[All Fields] AND “surveillance”[All Fields] AND “systems”[All Fields]) OR “communicable diseases, emerging”[MeSH Terms]) AND (“Internet search queries”[All Fields] OR “Google Trends”[All Fields] OR “Google Dengue Trends”[All Fields] OR “Google Flu Trends”[All Fields]) |
| Applied filters | None |
| Inclusion criteria | • Research articles where Internet search queries were applied as a data source for prediction modeling for epidemiological surveillance or early outbreak detection purposes/the association of GTD and the studied disease was assessed • Focused on emerging infectious diseases: HIV infections, SARS, COVID-19, Lyme disease, E. coli, Hantavirus, Dengue fever, West Nile virus, or Zika virus • Assessed the correlation between the Internet search queries/predicted cases and the actual cases In lower- to middle-income countries |
| Exclusion criteria | • Source of health information originated from non-Internet-based sources (e.g., electronic health records) or other Internet sources (e.g., social media) • Description of an algorithm, tool, or model in theory • Focus on non-communicable diseases, sexually transmitted infections (STIs), or pharmaceutical intervention use • Theoretical evaluation, only hypothetical mathematical modeling, calibration, etc. • Not original research article, e.g., content analysis, mathematical model, narrative analysis framework, conference articles, brief reports, letter to editors, communications, data articles, viewpoints, clinical studies, supplements, tutorials, reviews, or pilot studies • In high-income countries or in countries where Google is unavailable • Non-English publication |
Study selection process
The title and abstracts of all articles were reviewed independently by two authors (Beckhaus J and Belau MH). Conflicts in inclusion decisions were discussed and solved by consulting a third author (Becher H). In this study, emerging infectious diseases, as defined by the National Institute of Allergy and Infectious Diseases[22], were regarded. We therefore selected major emerging infectious diseases, namely HIV infections, SARS, Lyme disease, Escherichia coli O157:H7 (E. coli), Hantavirus, Dengue fever, West Nile virus, and the Zika virus, to compare a variety of pathogens and dissemination. We intend to provide a holistic picture of different processes and factors that may contribute to the applicability of Internet search queries data for these specific diseases. References were eligible for inclusion if they met the following criteria: (1) Internet search queries were applied as a data source for epidemiological surveillance or early outbreak detection; (2) focused emerging infectious diseases, i.e. HIV infections, SARS, COVID-19, Lyme disease, Dengue fever, E.coli, Hantavirus, West Nile virus, and Zika virus infection; (3) assessed the correlation between the Internet search queries/predicted cases and the actual cases; and (4) in lower- to middle-income countries. Exclusion criteria are explained in Table 1.
Data extraction
The following characteristics were extracted and summarized in duplicate by two authors (Beckhaus J and Belau MH) in a self-developed grid: author, year of publication, studied disease, study period, location searched, the purpose of the study, used keywords, used statistical methods (correlational analyses or regression analyses), and main findings and results. The respective correlation coefficients [Pearson’s correlation (r), Spearman’s correlation (ρ), and time lag correlation] were extracted.
Quality appraisal
References were critically appraised with a self-developed checklist, adapted from the methodological framework by Mavragani and Ochoa[18] and the review by Barros et al.[21]. The authors stated the importance of selection and explanation of the used region, keywords, period, and potential search categories from Google Trends data. Further, important characteristics for surveillance models as the type of system, data flow, resources, and data analysis were examined for the included studies[23]. Whether the article adequately discussed and whether a conflict of interest was stated were examined with the quality appraisal as well. Good, intermediate, and poor quality were defined as, respectively, no or one missing criterion, two to three missing criteria, and more than four missing criteria. The used checklists can be found in the Supplementary Materials.
RESULTS
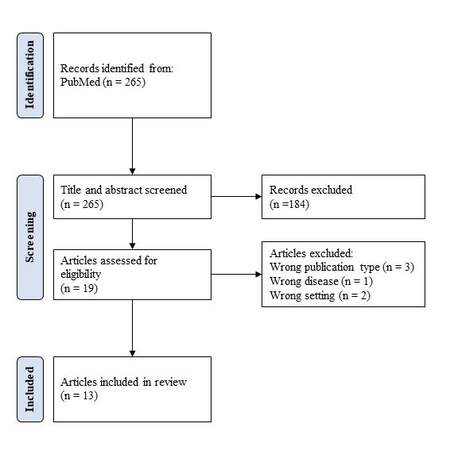
The search identified 265 relevant references [Figure 1]. The search date was 2 December 2021; 184 were excluded after the title and abstract screening. In total, 19 articles were eligible, and their full texts were screened. Six articles were excluded [Supplementary Material File 1]; three studies were not original research articles (e.g., the theoretical development of a mathematical model), one did not include an eligible disease, and two examined the wrong setting. Out of 13 included studies in the review, four focused on Dengue fever[25-28], four on Zika virus[29-32], and five on COVID-19 disease[33-37]. No study was identified that had met the inclusion criteria and focused on HIV infections, SARS, Lyme disease, E. coli, Hantavirus, or West Nile virus. It was striking that most of the 265 identified studies were from Western and high-income countries. In the studies included in our review, most were from Latin America (Brazil, Martinique, Honduras, El Salvador, Bolivia, Colombia, and Venezuela), three were from Asia (India and Indonesia), and Sub-Saharan Africa was underrepresented with only one study (South Africa). The study by Teng et al.[32] regarded the Zika virus globally.
Figure 1. Flow chart (According to Page et al.[24]).
Covid-19
Five included studies investigated COVID-19[33-37]. Schnoell et al.[36] and Sousa-Pinto et al.[37] regarded the early phase of the pandemic in different countries. Both studies assessed the correlation of weekly or average GTD with weekly new cases of COVID-19. For our comparison, only the results for Brazil and South Africa were selected. Schnoell et al.[36] used the keyword “coronavirus”; Sousa-Pinto et al.[37] included the symptoms of anosmia (loss of smell) and ageusia (loss of taste) in their trends analysis. The studied period was longer in the study by Schnoell et al.[36]: in Brazil, the period 26 February to 19 June 2020 and in South Africa from 6 March to 19 June 2020 was regarded. Sousa-Pinto et al.[37] assessed the early phase of the pandemic, from the first confirmed case in Brazil (25 February 2020) to 16 March 2020, which was no longer than three weeks in Brazil. Therefore, the study by Sousa-Pinto et al.[37] did not prove a significant correlation of GTD on anosmia and ageusia, two early symptoms with COVID-19 cases in Brazil during the early phase of the pandemic (r = -0.014, n.s.; r = -0.031, n.s.). The poor correlation may be related to the low number of cases of COVID-19 in Brazil during the regarded period. Therefore, they related the increase in the searched queries to media attention. Contrarily, Schnoell et al.[36] found a significant negative correlation of GTD with new COVID-19 cases in Brazil (ρ = -0.41, P < 0.001) and COVID-19 deaths (ρ = -0.42, P < 0.001). In South Africa, strong significant negative correlations were found of coronavirus-related GTD with new cases and deaths (ρ = -0.78, P < 0.001; ρ = -0.81, P < 0.001). The authors explained the negative values with the late onset of active cases in Brazil and South Africa. Aragón-Ayala et al.[33] examined the interest in COVID-19-related search queries (combined term) and daily cases of COVID-19 in Latin America and Caribbean countries (Peru, Panama, Colombia, Paraguay, Uruguay, Argentina, Bolivia, Ecuador, Costa Rica, Chile, Guatemala, Venezuela, Mexico, Dominican Republic, Brazil, Cuba, El Salvador, Honduras, Nicaragua, and Puerto Rico) from 30 December 2019 to 25 April 2020. They reported a high time-lag correlation of 0.72 (P < 0.001) for 18 out of 20 countries. The correlation was highest with a time lag of 5.67 days [Table 2]. In addition, Satpathy et al.[34] reported a high positive time-lag correlation between GT RSV (“COVID 19”, “COVID”, “social distancing”, “soap”, and “lockdown” at the national level: 0.68 (P < 0.01) (time lag of 11 days), 0.71 (P < 0.01) (time lag of 11 days), 0.62 (P < 0.01) (time lag of 11 days), 0.82 (P < 0.01) (time lag of 1 day), and 0.62 (P < 0.01) (time lag of 11 days)) and daily laboratory-confirmed COVID-19 cases in India. Seyyed Hosseini et al.[35] focused on Middle Eastern countries, where Syria was the only low- to middle-income country. In 2020, no significant Pearson’s correlation was found between search on GT and confirmed cases (r = 0.08; n.s.), but there was one for search on GT and COVID-19 deaths (0.17, P < 0.001). Therefore, the results for correlations between GTD and daily (or weekly) COVID-19 cases in different countries vary greatly.
Characteristics of included studies
| Ref. | Year | Studied disease | Location searched | Study period | Used keywords | Findings | Quality |
| Aragón-Ayala et al.[33] | 2021 | COVID-19 | Latin America | Dec 30 2019-Apr 25 2020 | Combined term of: “coronavirus + COVID-19 + SARS-Cov2 + nuevo coronavirus + 2019-nCoV” | No time lag correlation was found between this interest and national epidemiological indicators: ~0.72 (P < 0.001), 0.79 (P < 0.05) (time lag of 5.76 ± 13.35 days)a between RSV for COVID-19 and daily new cases | Good |
| Satpathy et al.[34] | 2021 | COVID-19 | India | Jan 1 2020-May 31 2020 | Coronavirus, corona, covid, covid 19, mask, sanitizer, social distancing, handwash, soap | GT RSV showed high time-lag correlation with both daily laboratory confirmed cases for the terms “COVID 19,” “COVID,” “social distancing,” “soap,” and “lockdown” at the national level: 0.68 (P < 0.01) (time lag of 11 days), 0.71 (P < 0.01) (time lag of 11 days), 0.62 (P < 0.01) (time lag of 11 days), 0.82 (P < 0.01) (time lag of 1 day), 0.62 (P < 0.01) (time lag of 11 days) | Good |
| Schnoell et al.[36] | 2021 | COVID-19 | Brazil; South Africa | Feb 26 2020-Jun 19 2020; Mar 06 2020-Jun 19 2020 | Coronavirus | Moderate to strong negative Spearman’s correlations of web-based interests and weekly covid-19 cases and deaths: -0.41 (P < 0.001) (cases), -0.42 (P < 0.001) (deaths); -0.78(P < 0.001) (cases), -0.81 (P < 0.001) (deaths) | Good |
| SeyyedHosseini et al.[35] | 2021 | COVID-19 | Syria | Jan 1 2020-Dec 31 2020 | Coronavirus + Corona virus + Corona + Covid19 + Covid سوریف + 19 نورک + دیفوک + دیفوک + انوروک + انوروک | No significant Pearson’s correlation was found between search on GT and confirmed cases but for search on GT and deaths for Syria: 0.08 (n.s.); 0.17 (P < 0.001) | Good |
| Sousa-Pinto et al.[37] | 2020 | COVID-19 | Brazil | first confirmed case of COVID-19-Mar 16 2020 | Anosmia, ageusia | COVID-19 related queries do not necessarily follow the evolution of the epidemic; for anosmia and ageusia, are more closely to media coverage; no significant Pearson’s correlation was found between average GT searches for anosmia/ageusia (topic), anosmia (disease) or ageusia (topic) and COVID-19 cases: -0.014 (n.s.), -0.031 (n.s.), 0.003 (n.s.) | Moderate |
| Chan et al.[27] | 2011 | Dengue fever | Bolivia, Brazil, India, Indonesia | 2003-2010 | Dengue-related Google search queries | models were able to adequately estimate true dengue acitivity according to official dengue case counts for the majority of the seasons during the time frame analyzed; high Pearson’s correlation between official dengue case count and dengue-related GT search queries: Bolivia 0.94, Brazil 0.92, India 0.87, Indonesia 0.9; holdout: Bolivia 0.83, Brazil 0.99, India 0.94, Indonesia 0.94 (no P-values reported) | Moderate |
| Husnayain et al.[26] | 2019 | Dengue fever | Indonesia | 2012-2016 | Disease definition, symptom, treatment, vector of disease | GTD had a linear time series pattern with official dengue report; GTD can be used for an early warning system because of time lag; novel tool before the increase of cases and during the outbreak; high Pearson’s correlation (time lag - 0) were found between dengue symptom, dengue, abbreviation of dengue and GTD: 0.937 (P < 0.05), 0.931 (P < 0.05), 0.921 (P < 0.05); season 2016: 0.954 (P < 0.05), 0.966 (P < 0.05), 0.950 (P < 0.05) | Good |
| Marques-Toledo et al.[28] | 2017 | Dengue fever | Brazil | Sep 2012-Oct 2016 | Dengue | Strong positive Pearso’s correlation of GTD and dengue cases in Brazil; useful for surveillance and prevention: 0.92 (P < 0.001) | Good |
| Monnaka et al.[25] | 2021 | Dengue fever | Brazil | Dec 31 2017-Mar 30 2019 | Dengue | Strong positive Pearson’s correlation between GTD and dengue cases in Sao Paulo: 0.91 (P < 0.05), 0.79 (P < 0.05) (time lag of - 1 week) | Good |
| Adebayo et al.[30] | 2017 | Zika virus | PAHO-region | May 2015-May 2016 | Zika virus[topic] | Strong Spearman’s correlation between online trend RSVs and number of suspected Zika cases; high cross-country variation: Brazil: 0.922 (P < 0.001); Colombia: 0.895 (P < 0.001); Martinique: 0.035 (n.s.); Honduras 0.748 (P < 0.001); El Salvador: 0.794 (P < 0.001) | Good |
| Morsy et al.[31] | 2018 | Zika virus | Brazil, Colombia | Jan 2016-July 2016; Aug 2015-May 2016 | Zika | Google search queries could be used to predict Zika cases 1 week earlier before the outbreak; auto-correlation of 0.986 (Brazil), 0.918 (Colombia) between observed and predicted Zika cases (no P-values reported) | Good |
| Strauss et al.[29] | 2020 | Zika virus | Venezuela | 2014-2016 | Zika | GTD was highly correlated with Zika at a time lag of + 1, with a Pearson’s correlation coefficient of 0.754 (no P-value reported) | Good |
| Teng et al.[32] | 2017 | Zika virus | globally | Feb 2016-Nov 2016 | Zika | GTD on Zika had statistically significant and positive Pearson’s correlations with the cumulative numbers of confirmed cases, suspected cases and total cases of ZIKV: 0.968 (P < 0.001), 0.980 (P < 0.001), 0.988 (P < 0.001) | Moderate |
Dengue fever
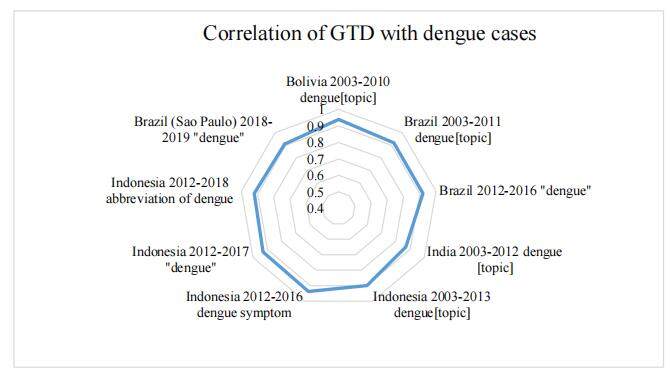
Four of the included thirteen studies examined the association of GTD and Dengue fever cases. The oldest included study[27], which was co-authored by Google employees, described the development of models that were able to estimate Dengue cases by using GTD. The regarded period was from 2003 to 2010, and higher-income countries were examined in addition to Bolivia, Brazil, India, and Indonesia. The found correlations ranged between r = 0.87 (India) and r = 0.94 (Brazil) for the overall dataset. In the dataset studied (which modeled an outbreak, irrespective of the calendar year), the correlation was lower in Bolivia (r = 0.83). The study did not provide information on the significance levels of the found correlation coefficients. In 2005, there was a major Dengue virus outbreak in a rural area of India (Kolkata, West Bengal) where using Google for Internet search was not yet common, compared to cities such as Delhi or Mumbai. The model on GTD therefore underestimated the cases, showing how dependent the fit is on Internet penetration. Nonetheless, the authors concluded that GTD can be used to adequately estimate true Dengue activity. A recently published study related to Dengue fever[26] examined the correlation between GTD and the Indonesian national surveillance report from 2012 to 2016. Three different trends were used, the Dengue symptoms, the term “Dengue”, and an Indonesian abbreviation of Dengue. Visually, the authors found a linear time-series pattern of the GTD with the official Dengue report. In contrast to other studies, Husnayain et al.[26] examined the overall period, included each year separately, and time lags of 1–3 weeks, respectively. When the Dengue symptoms were used for the GTD, the overall correlation was significantly high (r = 0.937, P < 0.05). In 2016, the correlation, therefore, was the highest for all years (r = 0.954, P < 0.05). With increasing time lag, the correlation decreased (time lag 0 weeks: r = 0.937, P < 0.05; time lag - 3 weeks: r = 0.517, P < 0.05). For the term “Dengue”, the overall correlation was significantly high (r = 0.93, P < 0.05), and again the highest for 2016 (r = 0.966, P < 0.05) and for zero time lag (r = 0.931, P < 0.05). When the Indonesian abbreviation of Dengue was used, the correlation was still high, but slightly lower compared to the previous keywords. Overall, the correlation was r = 0.921 (P < 0.05), higher in 2016 only (r = 0.95) and for zero time lag (r = 0.921, P < 0.05). The time lag analysis suggests that the correlation is the highest when no time lag is used. However, with one week’s time lag, the correlation remained high (r = 0.755-0.773). The authors concluded that GTD can be used for an early warning system and that the applied models can be implemented as a tool before or during a Dengue outbreak in Indonesia. Marques-Toledo et al.[28] investigated primarily Twitter data for Dengue fever surveillance in Brazil from September 2012 to October 2016 but used GTD for comparison. The used keyword was “Dengue”, which was significantly highly correlated with officially confirmed Dengue cases in the observed period (r = 0.92, P < 0.01). The authors concluded that GTD is a useful source for surveillance and prevention of Dengue fever outbreaks in Brazil. In addition, Monnaka et al.[25] assessed GTD and Dengue cases in Brazil but only for the region of Sao Paulo between 31 December 2017 and 30 March 2019. They found strong positive Pearson’s correlations of 0.91 (P < 0.05) and 0.79 (P < 0.05) with a time lag of - 1 week. The estimated correlation coefficients are summarized in a spider diagram [Figure 2]. In general, the reported values show a high correlation between GTD and Dengue fever cases, with the highest correlations in Bolivia from 2003 to 2010 (r = 0.94) and Indonesia from 2012 to 2016 (r = 0.937). The lowest correlation was found in India from 2003 to 2012 (r = 0.87). Indonesia was the most studied country in the included studies. The results are congruent since the values of the correlation coefficient were around 0.9. Nevertheless, the significance levels of the results were not included in the diagram and may therefore differ for each result.
Zika virus
The Zika virus was the topic of four included studies, one published in 2018 and two in 2017. Adebayo et al.[30] investigated the association of GTD and Zika virus infection incidence in the PAHO (Pan American health organization) region (e.g., Brazil, Colombia, Martinique, Honduras, and El Salvador) from May 2015 to May 2016. The reported Spearman’s correlation was high for Brazil (ρ = 0.922, P < 0.001), Colombia (ρ = 0.895, p < 0.001), Honduras (ρ = 0.748, P < 0.001), and El Salvador (ρ = 0.794, P < 0.001). For Martinique (ρ = 0.035, n.s.), no significant correlation was found. For Brazil, the authors performed an ARIMAX analysis to assess whether the press releases of WHO, Center for Disease Control and Prevention, and the Ministry of Health Brazil (MHB) were associated with GTD. Their results suggest that the online search trend of the previous week predicts the timing of the press releases by WHO and CDC but not MHB. They recommended using GTD as a complementation to traditional surveillance systems. Morsy et al.[31] examined whether the search query “Zika” could effectively predict Zika virus spread in Brazil (January-July 2016) and Colombia (August 2015 to May 2016). The observed auto-correlation coefficients were 0.986 and 0.918 between the observed and predicted Zika cases in Brazil and Colombia, respectively. They concluded that there was a good predictive capacity of the applied models. Therefore, with an applied time lag of one week, the performance of the model was still acceptable with the advance of time prediction, which is why the authors suggested using the time series regression model with a time lag of one week and autocorrelation. They concluded that Google search queries could predict Zika cases one week before the outbreak. A more general observation was made by Teng et al.[32] (2017), who explored the predictive power of GTD in the Zika spread worldwide from 12 February to 9 November 2016. They found a very highly significant Pearson’s correlation between GTD and confirmed Zika cases (r = 0.968, P < 0.001), suspected Zika cases (r = 0.98, P < 0.001), and total cases (r = 0.988, P < 0.001). Furthermore, they applied an ARIMA model which used the GTD as the exogenous variable to enhance the forecasting model. Hence, the applied ARIMA model showed good performance (compared to linear regression); it also predicted the number of Zika cases close to the actual observed cases in the early Zika epidemic in November 2016. The authors concluded that GTD is a useful data source, which can be included in prediction models. Strauss et al.[29] investigated whether GTD can be used to predict Zika cases in Venezuela. The regarded period was from 2014 to 2016; a high correlation of 0.754 was found, with a time lag of one week. They concluded that Google search queries represent a valuable and timely indicator for Dengue activity in Venezuela.
Quality of the included studies
The included studies were overall of moderate to good quality [Supplementary Material File 2]. Especially noticeable was the improvement of the study quality with later publication years. The latest publications were very detailed in the description of data entry concerning region, keyword, and period selection of GTD. The earliest publication[27] only provided brief information on the GTD itself, was not extensive on keyword selection, and explained the model building more in detail. The five most recent articles were overall of good quality[25,33-35]. In addition, Strauss et al.[29] implemented each quality criterion. Adebayo et al.[30], Morsy et al.[31], and Marques-Toledo et al.[28] fulfilled every criterion, except that there was not enough information on if and how data were edited. Chan et al.[27] explained in detail how data were edited, and excluded spurious spikes. The study by Husnayain et al.[26] was the only one that fulfilled all quality appraisal criteria and stated how they handled missing cases in their analysis (multiple imputation), which was not mentioned in any other included article. In addition, Schnoell et al.[36] explained how the GTD was normalized using the min-max method. Nonetheless, they did not provide enough information on the data analysis, so that another author would be able to reproduce their analysis. The same applies to the study by Sousa-Pinto et al.[37], where the presented method section about the data analysis would be insufficient for reproduction. Whether the data were edited was also not explained in detail. Lastly, the study by Teng et al.[32] was very brief in data entry, editing, and discussion. Clearly, a more comprehensive explanation of why a worldwide setting was selected and more comparisons with previous studies would have improved the quality of the article.
DISCUSSION
In the past five years, especially last year, there has been a growing usage of GTD in prediction models for epidemiological surveillance purposes. Our review aims to identify infodemiology studies to assess the use and applicability of GTD for infectious disease surveillance for low- to middle-income countries. Based on our findings, GTD in general appears to be a promising tool for prediction modeling of emerging infectious disease outbreaks or early warning systems.
GTD was used in five studies on COVID-19, four studies on Dengue, and four studies on Zika. All studies utilized GTD on disease-related keywords, either the disease itself or the disease definition, treatment, or symptoms. The assessed settings were in Latin and South America, India, Indonesia, South Africa, and Syria. In terms of the applicability, overall high correlations between GTD and the disease cases were found. For COVID-19, the study results are controversial. Schnoell[36], Sousa-Pinto[37], and Seyyed-Hosseini[35] found no or negative correlation, which might be due to the examined early phase of the pandemic, the great media attention[36,37], and the crisis and war in Syria[35]. Aragon-Ayala[33] and Satpathy[34] studied a longer period, later in the pandemic, and found a high correlation between GTD and COVID-19 cases. Regarding the short time frame, it is highly controversial to use Internet search queries during an emerging pandemic of a new disease. Conversely, Dengue fever is a well-known and widely spread disease. Dengue fever was discovered and studied before the Internet was widely accessible. The included studies considered already periods beginning from 2003, when the Internet penetration in the examined countries was still low. Nevertheless, a high correlation was reported over all the studies and different settings. GTD seems to be a suitable indicator for Dengue surveillance. Future studies should include time lag assessments to examine the predictive potential of GTD on Dengue surveillance. Regarding the Zika virus, GTD appeared applicable in surveillance. Since 2016, Zika virus outbreaks have occurred in many different countries and regions worldwide[38]. The regarded periods in the included studies started from 2014, when Internet penetration and technical developments were high. Overall, high correlations between GTD and cases were reported. The predictive value of GTD was presented, and a time lag of one week was consistent among the included studies that examined time lag correlations. For Dengue fever and Zika virus surveillance, GTD appears to be a useful indicator. In 2019, the fastest growth of subscribed mobile devices (with a majority of smartphones) was seen in Sub-Saharan Africa[39]. Since mobile devices and the Internet is accessible for the majority of the population, regardless of the economic situation of their country, more Internet search query data will become available. This can be an added value in public-health responses to outbreaks if a framework for data use is developed to ensure the systematic applicability of the data[39]. For COVID-19, the applicability of GTD in surveillance systems might increase in a couple of years, when Internet searches are more related to the actual infections than to media events. Evaluating a suitable time lag in the regression models is necessary for future studies. The nature of the disease and the dissemination should always be considered, especially in the case of COVID-19 where we know that one can already be infectious without having symptoms or even noticing the disease. The Internet search behavior of the individual cannot function as an early predictor of the disease. Therefore, more infodemiological studies on COVID-19 are needed to allow a robust conclusion on the applicability of GTD.
There are several limitations to our review. The search was limited to one database, and no additional hand search or reference list search was added to the search strategy. Our review might have missed scientific articles from other disciplines than medical research. We did not use setting specific databases (e.g., LILACS or African Journal Online) for our literature research. The language was restricted to English, and we therefore might have missed relevant articles in other languages that regarded low- to middle-income countries. Hence, we did not find any studies investigating African countries. The quality of the included studies was assessed subjectively, and the self-developed tool is not operationalized or evaluated yet. Despite its limitations, this review has some major strengths, including the performed systematic search in Medline, with relevant search terms identified through recent literature. Since the focus was to investigate the use and applicability of GTD for epidemiological surveillance in low- to middle-income countries, it adds another insight and reveals the need for further research regarding these settings. The included quality appraisal added the necessity of standard requirements in infodemiology studies. All the included studies were ecological studies, focusing on Internet search queries and their correlation with infectious disease cases. The included studies were sufficient to show that GTD is used and applicable in Dengue and Zika surveillance. Using GTD should not replace traditional surveillance; rather, it is meant to supplement it when there is insufficient surveillance because of missing data sources. Besides, we included only studies that used GTD specifically. There might have been other useful Internet search query data utilized by studies and applied in surveillance. Therefore, this might have induced selection bias. Additionally, Chinese studies were not included because Google is not available there.
Compared to previous reviews on web-based infectious disease surveillance or infodemiology studies, this review revealed the same opportunities and challenges of using GTD. For example, Choi et al.[20] evaluated 11 surveillance systems that used online sources, four of which used Internet search queries. They concluded that the advantages of those systems were the low costs and their intuitive and near the real-time adaptable operation. Conversely, they explained the disadvantages of inaccuracy, false predictions, and potential violation of privacy. Milinovich et al.[6] regarded Internet-based surveillance systems for monitoring emerging infectious diseases, where they also revealed the focus on high-income countries. With regard to the emergence of infectious diseases all over the world, the authors suggested investigating the global surveillance on Internet-based resources further, as Teng et al.[32] did already for the Zika virus. Nonetheless, surveillance systems on GTD do not provide the same capacity as traditional systems and can rather supplement traditional systems[6]. Gianfredi et al.[1] discussed the issue of the unmeasured influence on the search behavior of the public. In times of rapid changing always-available online news, when messages spread fast, the search behavior can be more influenced by media than by actual events. Non-evidence-based rumors and false beliefs might influence the search behavior and the resulting trends, which can have consequences for decision-making in public health[1]. Barros et al.[21] underlined this effect, stating that media events significantly affect the reliability of search queries data. The majority of the included studies in their recent systematic review used search queries (and social media) as Internet-based sources. They also underlined the non-generalizability of the data, since the Internet users might be higher educated and younger than the general public. The greatest limitation according to Barros et al.[21] is the lack of transparency on how the data are obtained by Google.
GTD in epidemiological research can be useful as an additional data source to complement current disease surveillance systems. It should not be considered as the only data source, and interpretations and implementations should be handled with caution. If more research is done on the methodology of the implementation of GTD in the real-world setting, better and more precise implications can be withdrawn.
CONCLUSION
The low costs and availability of GTD remain promising for supplementing future surveillance systems in low- to middle-income countries, but there is an urgent need for a standard methodological framework for the use and application of GTD. The implementation of GTD in surveillance systems for Dengue fever and Zika can be recommended to complement existing systems. The connection of Internet search queries with zoonosis surveillance builds bridges between environmental and human health. High-quality infodemiology studies from and on low- to middle-income settings are needed to receive more relevant and applicable results in these regions.
DECLARATIONS
AcknowledgementsThis work has been done within the ESIDA (Epidemiological Surveillance for Infectious Diseases in Sub-Sahara Africa) project, funded by the German Federal Ministry of Education and Research.
Authors’ contributionsMade substantial contributions to conception and design of the study, performed data acquisition, wrote the manuscript and performed data analysis and interpretation: Julia Beckhaus
Contributed to writing the manuscript, provided administrative support: Heiko Becher
Made substantial contributions to conception and design of the study, supervised the research and contributed to writing the manuscript: Matthias Hans Belau
Availability of data and materialsNot applicable.
Financial support and sponsorshipThe research was supported by the German Federal Ministry of Education and Research, GRANT number 01DU20005C, ESIDA.
Conflicts of interestAll authors declared that there are no conflicts of interest.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2022.
REFERENCES
1. Gianfredi V, Bragazzi NL, Nucci D, et al. Harnessing big data for communicable tropical and sub-tropical disorders: implications from a systematic review of the literature. Front Public Health 2018;6:90.
3. World Health Organization (WHO). The top 10 causes of death. Available from: https://www.who.int/news-room/fact-sheets/detail/the-top-10-causes-of-death [Last accessed on 22 Mar 2022].
4. Bönecke, J. ESIDA: Epidemiological surveillance for infectious diseases in sub-saharan africa. Available from: https://www.haw-hamburg.de/en/research/projects-a-z/research-projects/project/project/show/esida/ [Last accessed on 22 Mar 2022].
5. . Cookson, ST, Buehler JW. Emergency and disaster health surveillance. In: Ahrens W. Pigeot I, editors. Handbook of Epidemiology. New York: Springer; 2014. p. 731-59.
6. Milinovich GJ, Williams GM, Clements ACA, Hu W. Internet-based surveillance systems for monitoring emerging infectious diseases. Lancet Infect Dis 2014;14:160-8.
7. Riel N, Auwerx K, Debbaut P, Van Hees S, Schoenmakers B. The effect of Dr Google on doctor-patient encounters in primary care: a quantitative, observational, cross-sectional study. BJGP Open 2017;1:bjgpopen17X100833.
8. Samaras L, García-Barriocanal E, Sicilia MA. Comparing Social media and Google to detect and predict severe epidemics. Sci Rep 2020;10:4747.
9. Google Trends. The year in review. Available from: https://trends.google.com/trends/yis/2020/GLOBAL/ [Last accessed on 22 Mar 2022].
10. Cousins HC, Cousins CC, Harris A, Pasquale LR. Regional infoveillance of COVID-19 case rates: analysis of search-engine query patterns. J Med Internet Res 2020;22:e19483.
11. Effenberger M, Kronbichler A, Shin JI, Mayer G, Tilg H, Perco P. Association of the COVID-19 pandemic with Internet Search Volumes: a Google TRENDSTM Analysis. Int J Infect Dis 2020;95:192-7.
12. Higgins TS, Wu AW, Sharma D, Illing EA, Rubel K, Ting JY. Snot Force Alliance. Correlations of Online Search Engine Trends With Coronavirus Disease (COVID-19) Incidence: infodemiology study. JMIR Public Health Surveill 2020;6:e19702.
13. Ginsberg J, Mohebbi MH, Patel RS, Brammer L, Smolinski MS, Brilliant L. Detecting influenza epidemics using search engine query data. Nature 2009;457:1012-4.
14. Kandula S, Shaman J. Reappraising the utility of Google flu trends. PLoS Comput Biol 2019;15:e1007258.
15. Lazer D, Kennedy R, King G, Vespignani A. Big data. The parable of Google Flu: traps in big data analysis. Science 2014;343:1203-5.
16. Nichols JA, Herbert Chan HW, Baker MAB. Machine learning: applications of artificial intelligence to imaging and diagnosis. Biophys Rev 2019;11:111-8.
17. Eysenbach G. Infodemiology and infoveillance: framework for an emerging set of public health informatics methods to analyze search, communication and publication behavior on the Internet. J Med Internet Res 2009;11:e11.
18. Mavragani A, Ochoa G. Google trends in infodemiology and infoveillance: methodology framework. JMIR Public Health Surveill 2019;5:e13439.
20. Choi J, Cho Y, Shim E, Woo H. Web-based infectious disease surveillance systems and public health perspectives: a systematic review. BMC Public Health 2016;16:1238.
21. Barros JM, Duggan J, Rebholz-Schuhmann D. The application of internet-based sources for public health surveillance (infoveillance): systematic review. J Med Internet Res 2020;22:e13680.
22. National Institute of Allergy and Infectious Diseases (NIAID). NIAID emerging infectious diseases / pathogens. Available from: https://www.niaid.nih.gov/research/emerging-infectious-diseases-pathogens [Last accessed on 22 Mar 2022].
23. Nsubuga P, White ME, Thacker SB, et al. Public health surveillance: a tool for targeting and monitoring interventions. In: Jamison DT, Breman JG, Measham AR, et al., editors. Disease Control Priorities in Developing Countries. 2nd edition. USA: Washington (DC): The International Bank for Reconstruction and Development / The World Bank; 2006. Chapter 53.
24. Page MJ, McKenzie JE, Bossuyt PM, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 2021;372:n71.
25. Monnaka VU, Oliveira CAC. Google trends correlation and sensitivity for outbreaks of dengue and yellow fever in the state of São Paulo. Einstein (Sao Paulo) 2021;19:eAO5969.
26. Husnayain A, Fuad A, Lazuardi L. Correlation between Google Trends on dengue fever and national surveillance report in Indonesia. Glob Health Action 2019;12:1552652.
27. Chan EH, Sahai V, Conrad C, Brownstein JS. Using web search query data to monitor dengue epidemics: a new model for neglected tropical disease surveillance. PLoS Negl Trop Dis 2011;5:e1206.
28. Marques-Toledo CA, Degener CM, Vinhal L, et al. Dengue prediction by the web: Tweets are a useful tool for estimating and forecasting Dengue at country and city level. PLoS Negl Trop Dis 2017;11:e0005729.
29. Strauss R, Lorenz E, Kristensen K, et al. Investigating the utility of Google trends for Zika and Chikungunya surveillance in Venezuela. BMC Public Health 2020;20:947.
30. Adebayo G, Neumark Y, Gesser-Edelsburg A, Abu Ahmad W, Levine H. Zika pandemic online trends, incidence and health risk communication: a time trend study. BMJ Glob Health 2017;2:e000296.
31. Morsy S, Dang TN, Kamel MG, et al. Prediction of Zika-confirmed cases in Brazil and Colombia using Google Trends. Epidemiol Infect 2018;146:1625-7.
32. Teng Y, Bi D, Xie G, et al. Dynamic forecasting of Zika epidemics using google trends. PLoS One 2017;12:e0165085.
33. Aragón-Ayala CJ, Copa-Uscamayta J, Herrera L, Zela-Coila F, Quispe-Juli CU. Interest in COVID-19 in Latin America and the Caribbean: an infodemiological study using Google Trends. Cad Saude Publica 2021;37:e00270720.
34. Satpathy P, Kumar S, Prasad P. Suitability of Google Trends™ for digital surveillance during ongoing COVID-19 epidemic: a case study from India. Disaster Med Public Health Prep 2021:1-10.
35. SeyyedHosseini S, BasirianJahromi R. COVID-19 pandemic in the Middle East countries: coronavirus-seeking behavior versus coronavirus-related publications. Scientometrics 2021:1-21.
36. Schnoell J, Besser G, Jank BJ, et al. The association between COVID-19 cases and deaths and web-based public inquiries. Infect Dis (Lond) 2021;53:176-83.
37. Sousa-Pinto B, Anto A, Czarlewski W, Anto JM, Fonseca JA, Bousquet J. Assessment of the impact of media coverage on COVID-19-related google trends data: infodemiology study. J Med Internet Res 2020;22:e19611.
38. Vue D, Tang Q. Zika virus overview: transmission, origin, pathogenesis, animal model and diagnosis. Zoonoses (Burlingt) 2021:1.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Beckhaus J, Becher H, Belau MH. The use and applicability of Internet search queries for infectious disease surveillance in low- to middle-income countries. One Health Implement Res 2022;2:15-28. http://dx.doi.org/10.20517/ohir.2022.01
AMA Style
Beckhaus J, Becher H, Belau MH. The use and applicability of Internet search queries for infectious disease surveillance in low- to middle-income countries. One Health & Implementation Research. 2022; 2(1): 15-28. http://dx.doi.org/10.20517/ohir.2022.01
Chicago/Turabian Style
Beckhaus, Julia, Heiko Becher, Matthias Hans Belau. 2022. "The use and applicability of Internet search queries for infectious disease surveillance in low- to middle-income countries" One Health & Implementation Research. 2, no.1: 15-28. http://dx.doi.org/10.20517/ohir.2022.01
ACS Style
Beckhaus, J.; Becher H.; Belau MH. The use and applicability of Internet search queries for infectious disease surveillance in low- to middle-income countries. One. Health Implement. Res. 2022, 2, 15-28. http://dx.doi.org/10.20517/ohir.2022.01
About This Article
Copyright
Data & Comments
Data

 Cite This Article 10 clicks
Cite This Article 10 clicks










Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.